Risk of severe clinical outcomes among persons with SARS-CoV-2 infection with differing levels of vaccination during widespread Omicron (B.1.1.529) and Delta (B.1.617.2) variant circulation in Northern California: A retrospective cohort study -

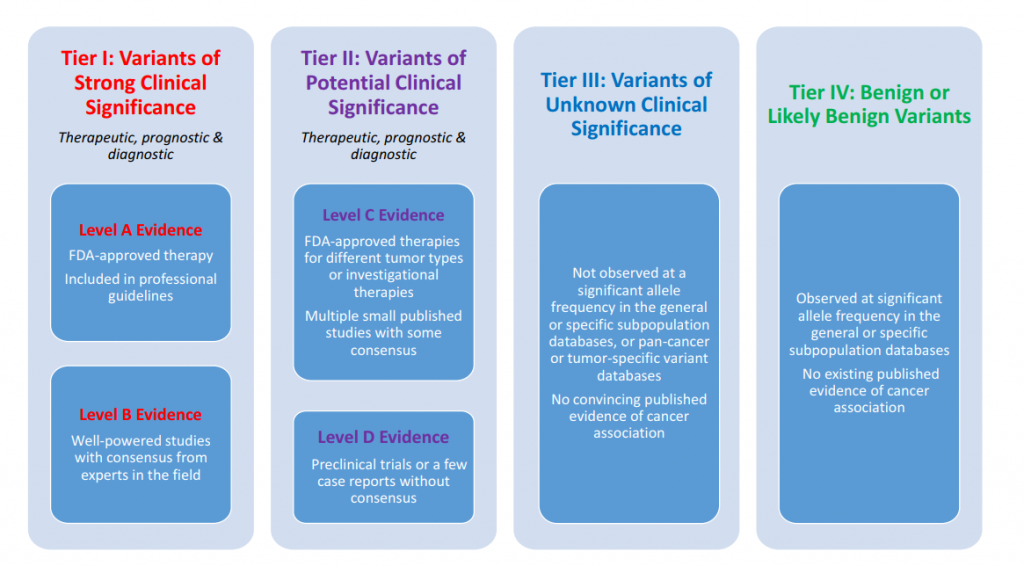
Standards for the classification of pathogenicity of somatic variants in cancer (oncogenicity): Joint recommendations of Clinical Genome Resource (ClinGen), Cancer Genomics Consortium (CGC), and Variant Interpretation for Cancer Consortium (VICC ...

Clinical Variant Classification: A Comparison of Public Databases and a Commercial Testing Laboratory. | Semantic Scholar

The Clinical Variant Analysis Tool: Analyzing the evidence supporting reported genomic variation in clinical practice - ScienceDirect
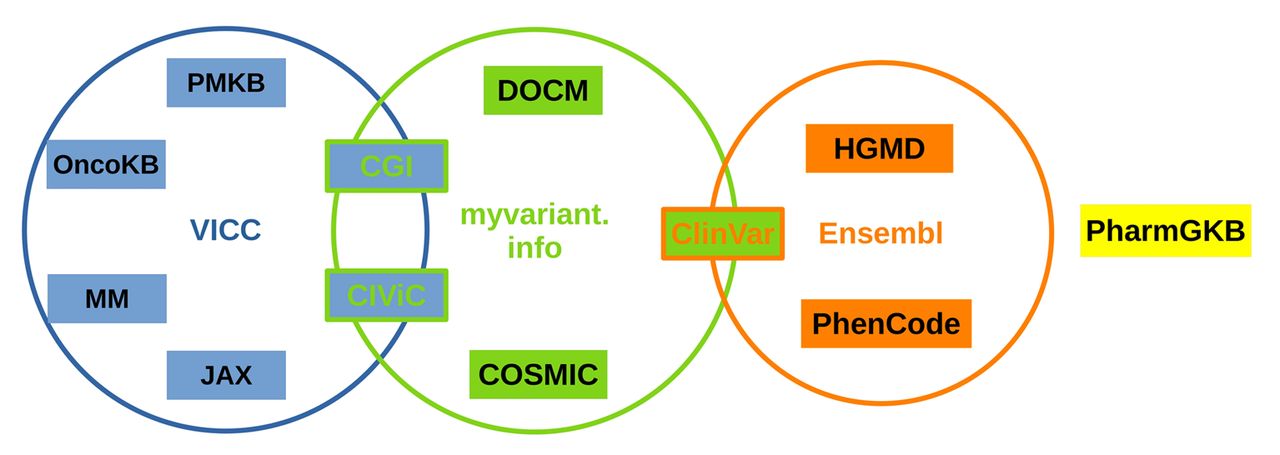
Comparison of Open-access Databases for Clinical Variant Interpretation in Cancer: A Case Study of MDS/AML | Cancer Genomics & Proteomics
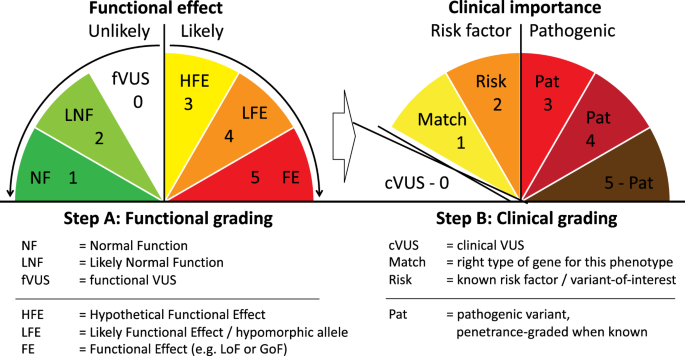
Stepwise ABC system for classification of any type of genetic variant | European Journal of Human Genetics

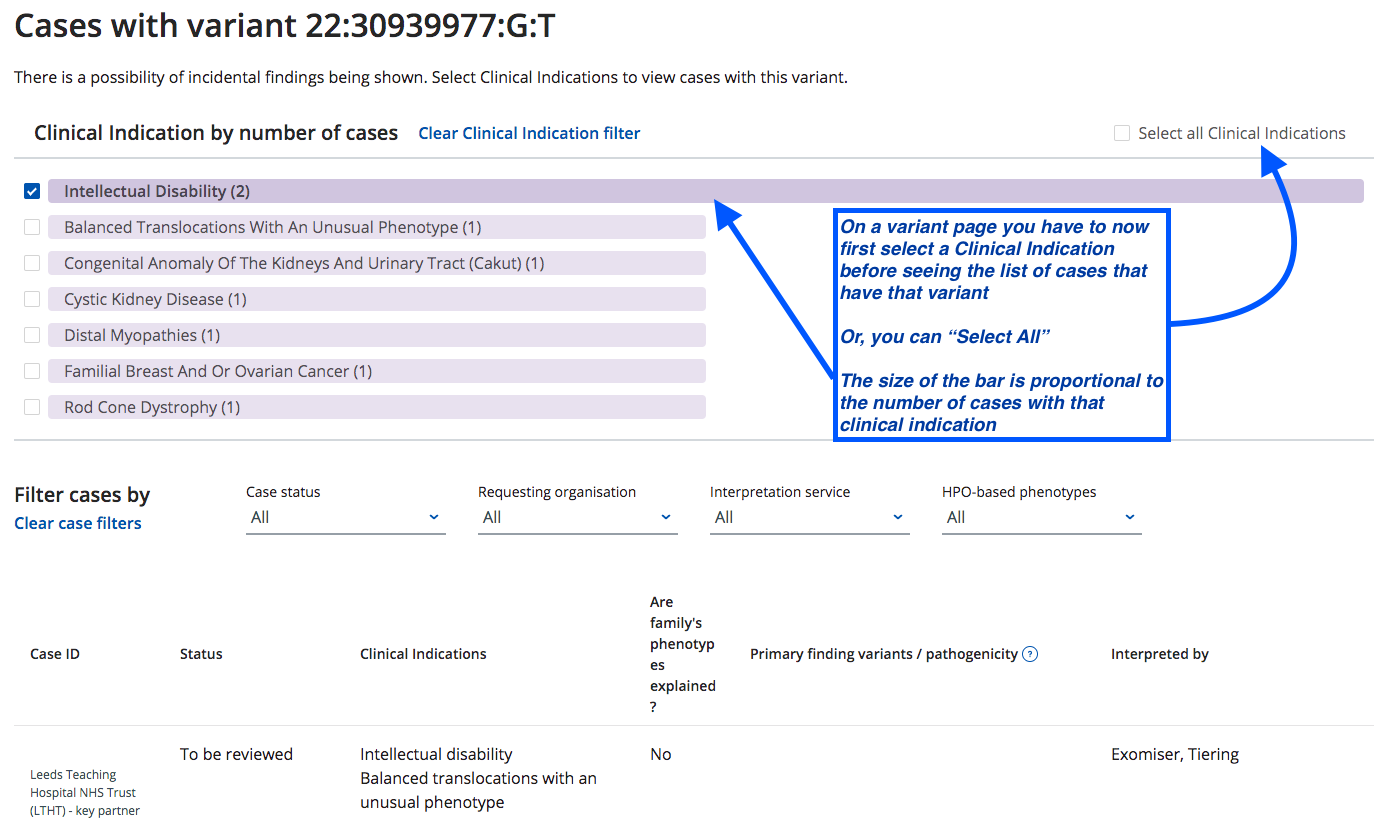
Clinical information can lead to finding a variant that might otherwise be missed - Blueprint Genetics
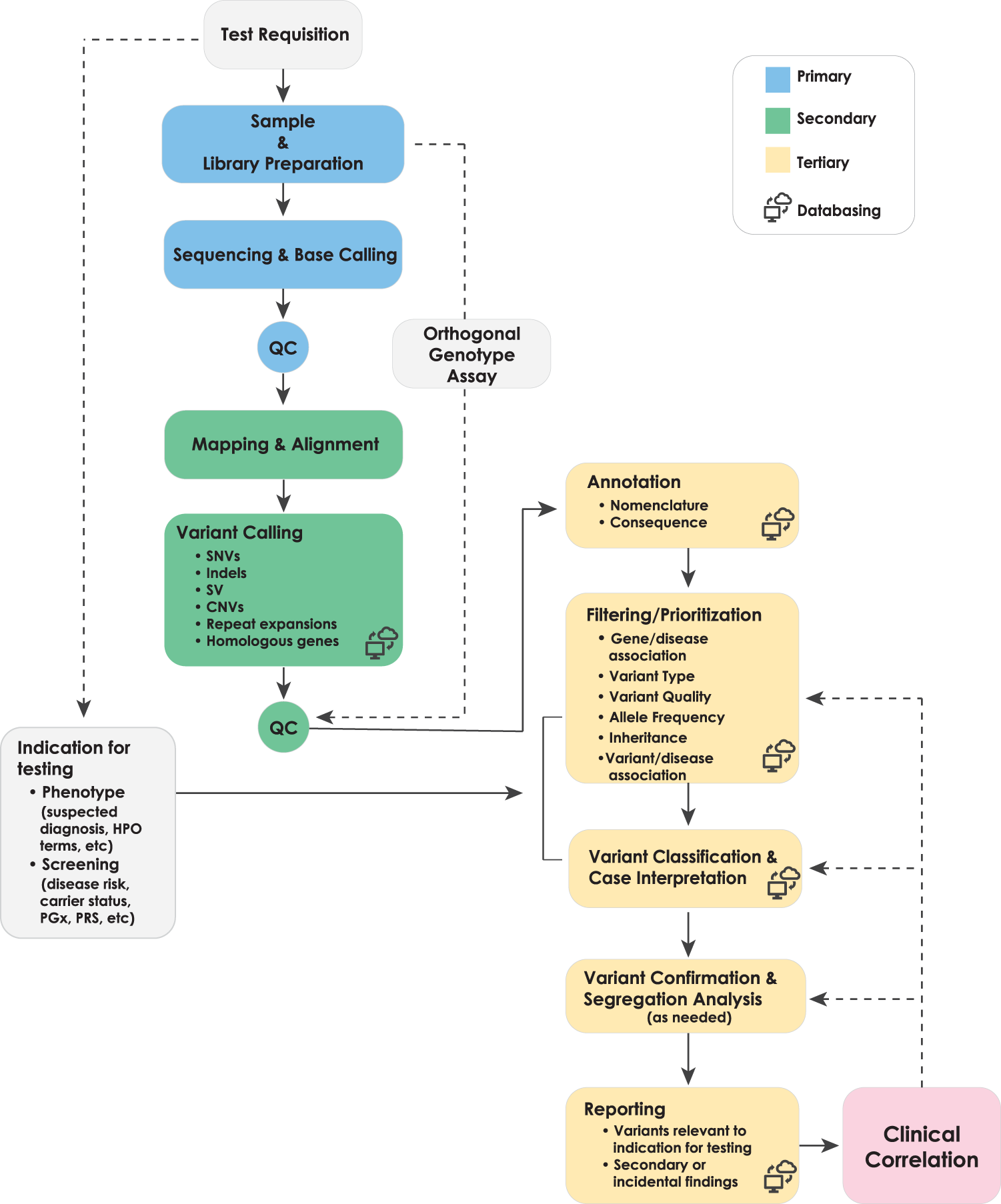
Best practices for the interpretation and reporting of clinical whole genome sequencing | npj Genomic Medicine

The Clinical Variant Analysis Tool: Analyzing the evidence supporting reported genomic variation in clinical practice - ScienceDirect

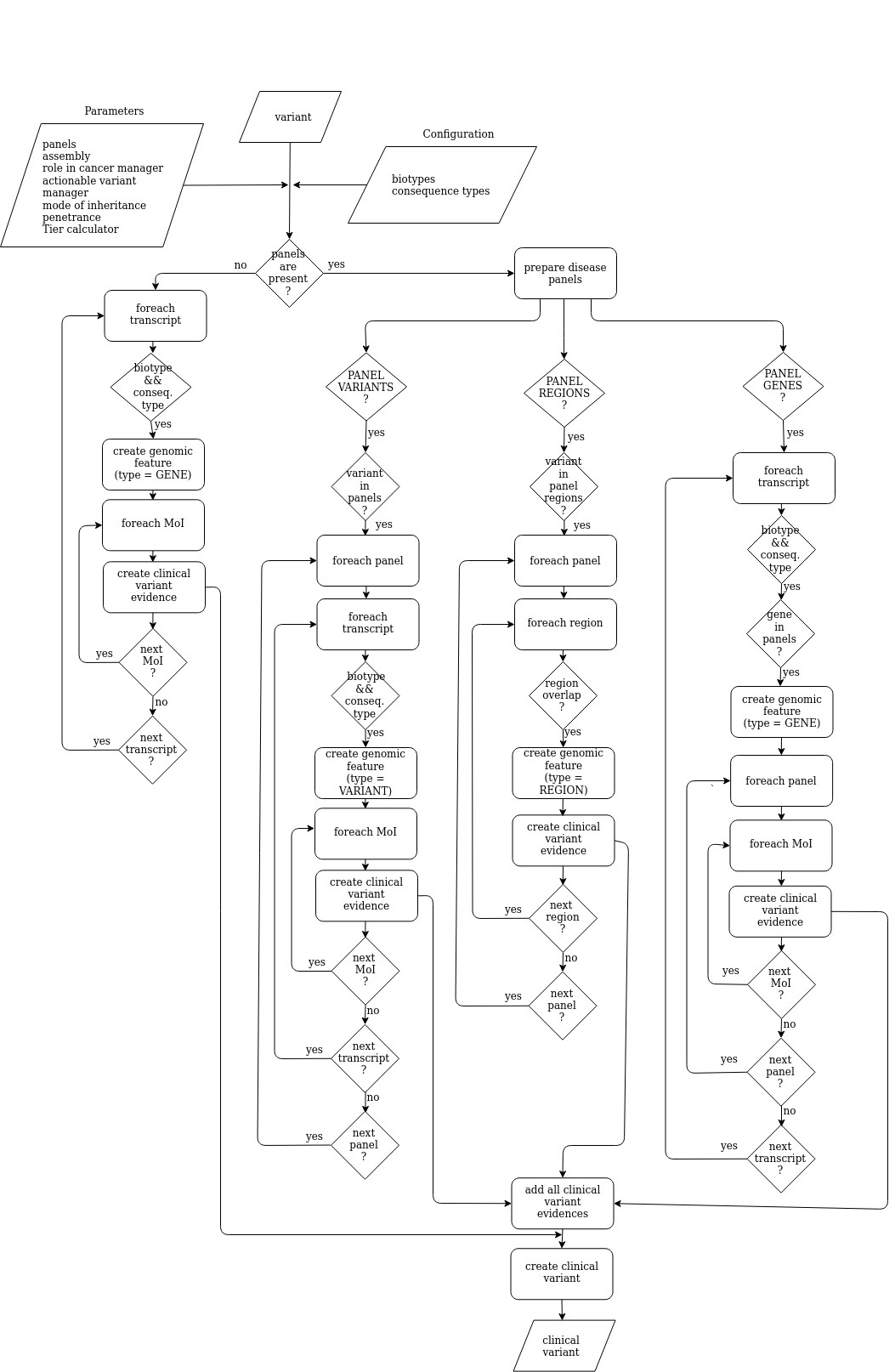
Schematic view of the clinical variant interpretation process. In a... | Download Scientific Diagram

Nothing's for sure, that's for sure: Evaluating variants of uncertain significance | Beyond the Ion Channel
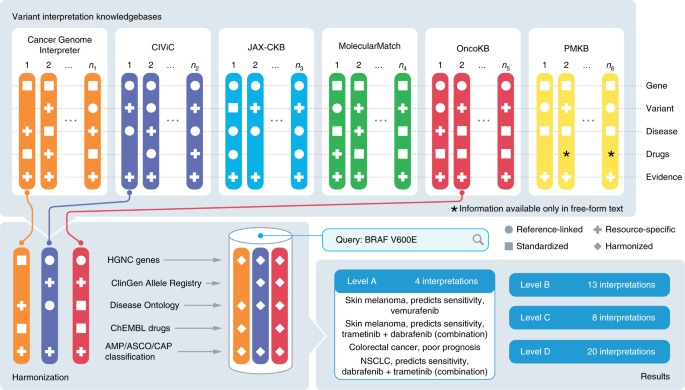
A harmonized meta-knowledgebase of clinical interpretations of somatic genomic variants in cancer | Nature Genetics

Assessing Variants in a Known Gene: Clinical Variant Classification and Use of the ClinGen Variant Curation Interface - ClinGen | Clinical Genome Resource